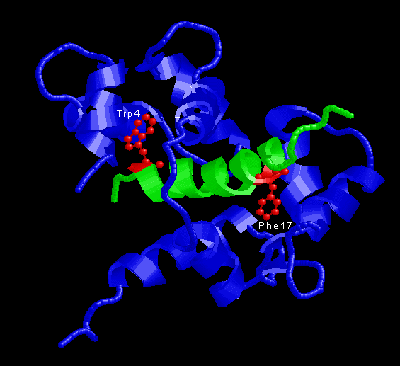
In the picture above, the two key residues are in red, M13 in green and calmodulin in blue.
PDB ID: 2BBM
Medline Entry
PDB file
Link to Calmodulin-binding Target Database
M13 peptide is classified under 1-14 motif.
Biology
Myosin
Myosin is an important motor molecule in muscle cells. Its activity requires actin.Myosin is composed of the heavy chain and the light chain. Heavy chain contains ATPase domain which hydrolyses ATP and use the energy to change the conformation and generate force. Light chain is at the neck region of myosin, and often is used as a regulatory domain. In vertebrates, phosphorylation of myosin light chain activates myosin.
M13
M13 is a synthetic peptide. Its sequence is the same as the calmodulin-binding domain of skeletal muscle myosin light chain kinase (skMLCK) (residues 577-602). It was used to demonstrate that calmodulin can bind the peptide without the presence of the entire protein.Calmodulin
Calmodulin is a ubiquitous protein that regulates many cellular processes by monitoring the calcium level in a cell. Calmodulin regulates cellular and physiological processes by interacting with various target enzymes such as myosin light chain kinase (MLCK), calmodulin-dependent kinases, protein phosphatase calcineurin, phosphodiesterase, nitric oxide synthase, Ca2+-ATPase pumps, as well as cytoskeletal structural proteins.How they work
When a muscle cell receives a signal from the neurons, calcium ions are released from sarcoplasmic reticulum (SR) which normally stores calcium ions. Calmodulin, normally present in the cytoplasm, binds calcium ions and changes conformation. This change in conformation allows calmodulin to bind many different target proteins, one of which is Myosin Light Chain Kinase (MLCK). When MLCK is bound to calmodulin, it is activated, and thus phosphorylates Myosin Light Chain. When myosin light chain is phosphorylated, myosin is allowed to hydrolyse ATP and use the energy to go through the myosin cross-bridge cycle.Structure
The conformation of the EF-hand domain stays relatively constant before and after binding to a peptide. However, the backbone linker region unravels in the middle to allow bending of the backbone. This bending allows peptides of limited but various length to bind calmodulin.Key features to note from the structure are the alpha-helical conformation of the M13 peptide and the two bulky hydrophobic residues, Trp4 and Phe17 of the M13. Trp4 and Phe17 make extensive contacts with calmodulin at the deep hydrophobic pockets on both domains of calmodulin. For this reason, Trp4 and Phy17 are considered to be the anchor residues for this interaction.
Also, note the orientation of the peptide relative to calmodulin. The
C-terminal domain of calmodulin interacts with the N-terminal end of the
peptide while the N-terminal domain of calmodulin interacts with the C-terminal
end of the peptide. The speculated reason for this particular orientation
is that the C-terminal domain of calmodulin is more negatively charged,
and the N-terminal end of the peptide is more positively charged. Therefore,
this orientation is preferred over the parallel orientation for electrostatic
reasons.